Project Overview
The goal of this project is to characterize iron signaling pathways in yeast model systems using structural, biophysical, and genetic methods to probe the functional and physical interactions that govern these signaling pathways.
The unique chemical properties of iron make it an essential but potentially toxic
protein cofactor. Iron is capable of performing redox chemistry that is required for
basic cellular functions (e.g. electron transfer, catalysis, oxygen transport), yet
this same property can lead to the production of reactive oxygen species that damage
cellular components. Organisms must maintain iron levels at sufficient, non-toxic
levels for survival.
Defects in iron metabolism are common disorders that lead to anemias, Friedreich’s
ataxia, muscles myopathies, mitochondrial dysfunction disorders, and hemochromatosis.
In fact, due to the low bioavailability of iron, iron deficiency is the most common
and widespread nutritional disorder in the world. Significant progress in identifying
specific proteins involved in iron metabolism has provided a glimpse at the elaborate
control mechanisms required to regulate this essential metal and revealed critical
insight into the pathophysiology of iron-associated genetic disorders. However, there
are substantial gaps in our fundamental understanding of iron homeostasis mechanisms
at both the cellular and molecular level that require further study. Filling these
gaps will be essential for preventing and treating disorders of iron metabolism.
Yeast are simple, single-cell eukaryotes that are inexpensive and easy to maintain and genetically manipulate in the lab, yet express many of the same iron homeostasis systems as human cells. In particular, most of the iron-binding yeast proteins we study have close human homologues. Thus uncovering their specific functional interactions in yeast will provide critical clues about the roles of these proteins in human metabolism. Another benefit of using yeast model organisms like Saccharoymces cerevisiae and Schizosaccharomyce pombe is that they are non-toxic to humans. For example, S. cerevisiae is also known as bakers' or brewers' yeast used for making bread, beer, and wine.
Iron Regulation in S. cerevisiae
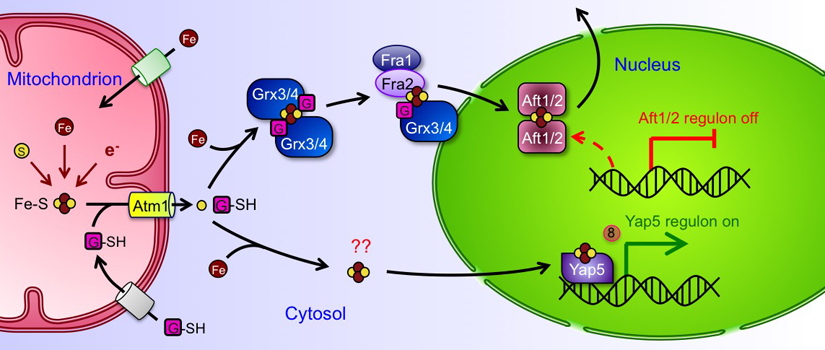
Regulation of iron metabolism in S. cerevisiae is controlled at the transcriptional level by low (Aft1 and Aft2) and high iron sensing (Yap5) transcriptional activators. When iron levels drop below a critical threshold, Aft1 and its paralogue Aft2 activate expression of proteins required for iron uptake at the plasma membrane, vacuolar iron transport, and mitochondrial iron metabolism. The net result is mobilization of iron from extracellular sources and intracellular stores. During iron sufficiency, Aft1 and Aft2 are inactivated while Yap5 turns on expression of genes involved in sequestering excess iron to protect the cell from iron toxicity.
Research Progress by Our Group
Although much is known about Aft1/2 and Yap5 target genes and the mechanisms for iron uptake, storage, and trafficking in yeast, less is known about the molecular mechanisms for controlling Aft1/2 and Yap5 activity and the specific iron signals to which Aft1/2 and Yap5 respond. Our research is focused on addressing these issues. We and our collaborators established that the monothiol glutaredoxins Grx3 and Grx4, the BolA-like protein Fra2 and the aminopeptidase P-like protein Fra1 function together in an Fe-S cluster-dependent signaling pathway that inhibits the activity of Aft1 and Aft2 during iron sufficiency. We showed that Grx3/4 and Fra2 form GSH-ligated, [2Fe-2S]2+-bridged heterodimers with unusual Fe-S coordination chemistry. Moreover, we solved the long-standing puzzle of the Aft1/2 regulation mechanism by demonstrating that Fra2-Grx3 transfers a [2Fe-2S] cluster to conserved cysteines in Aft2, which drives Aft2 dimerization, DNA dissociation, and nuclear export, with the net effect of turning off iron uptake and storage genes. Finally, we have shown that the human homologues of Grx3/4 and Fra2 form [2Fe-2S]-bridged complexes with analogous cluster coordination environments, suggesting a conservation of structure and function for these protein families across evolution. Numerous other researchers have built on our findings in S. cerevisiae to better define the roles of Grxs and BolA proteins in bacteria, protist, yeast, plant, and human iron metabolism. Overall, our studies have uncovered dedicated pathways for Fe-S cluster delivery in the cell and highlighted the importance of Fe-S clusters as key sensory molecules for monitoring intracellular iron conditions.